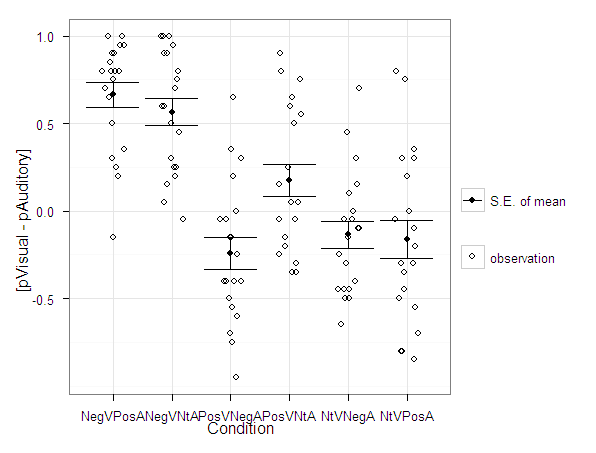
Over at the stats.se site I have come across a few questions demonstrating the power of utilizing dot plots to visualize experimental results.
- Improving data analysis through a better visualization of data?
- Analyze and visualize participants response towards particular condition
- Alternative graphics to handle bar plots
- How to avoid overplotting (for points) using base-graph? from Stackoverflow
Also some interesting discussion on what error bars to plot in similar experiments is in this question, Follow up: In a mixed within-between ANOVA plot estimated SEs or actual SEs?
Here I will give two examples utilizing SPSS and R to produce similar plots. I haven’t annotated the code that much, but if you need anything clarified on what the code is doing let me know in the comments. The data is taken from this question on the stats site.
Citations of Interest to the Topic
- Drummond, Gordon B. & Sarah L. Vowler. 2011. Show the data, don’t conceal them. The Journal of Physiology 598(8): 1861-1863. PDF available from publisher.
- Koyama, Tatsuki. Beware of Dynamite
SPSS Code to generate below dot plot
*******************************************************************************************. data list free /NegVPosA NegVNtA PosVNegA PosVNtA NtVNegA NtVPosA.
begin data
0.5 0.5 -0.4 0.8 -0.45 -0.3
0.25 0.7 -0.05 -0.35 0.7 0.75
0.8 0.75 0.65 0.9 -0.15 0
0.8 0.9 -0.95 -0.05 -0.1 -0.05
0.9 1 -0.15 -0.35 0.1 -0.85
0.8 0.8 0.35 0.75 -0.05 -0.2
0.95 0.25 -0.55 -0.3 0.15 0.3
1 1 0.3 0.65 -0.25 0.35
0.65 1 -0.4 0.25 0.3 -0.8
-0.15 0.05 -0.75 -0.15 -0.45 -0.1
0.3 0.6 -0.7 -0.2 -0.5 -0.8
0.85 0.45 0.2 -0.05 -0.45 -0.5
0.35 0.2 -0.6 -0.05 -0.3 -0.35
0.95 0.95 -0.4 0.55 -0.1 0.8
0.75 0.3 -0.05 -0.25 0.45 -0.45
1 0.9 0 0.5 -0.4 0.2
0.9 0.25 -0.25 0.15 -0.65 -0.7
0.7 0.6 -0.15 0.05 0 -0.3
0.8 0.15 -0.4 0.6 -0.05 -0.55
0.2 -0.05 -0.5 0.05 -0.5 0.3
end data.
dataset name dynamite.
*reshaping the data wide to long, to use conditions as factors in the plot.
varstocases
/make condition_score from NegVPosA to NtVPosA
/INDEX = condition (condition_score).
*dot plot, used dodge symmetric instead of jitter.
GGRAPH
/GRAPHDATASET dataset = dynamite NAME="graphdataset" VARIABLES=condition condition_score MISSING=LISTWISE
REPORTMISSING=NO
/GRAPHSPEC SOURCE=INLINE.
BEGIN GPL
SOURCE: s=userSource(id("graphdataset"))
DATA: condition=col(source(s), name("condition"), unit.category())
DATA: condition_score=col(source(s), name("condition_score"))
GUIDE: axis(dim(1), label("condition"))
GUIDE: axis(dim(2), label("condition_score"))
ELEMENT: point.dodge.symmetric(position(condition*condition_score))
END GPL.
*confidence interval plot.
*cant get gpl working (maybe it is because older version) - will capture std error of mean.
dataset declare mean.
OMS /IF LABELS = 'Report'
/DESTINATION FORMAT = SAV OUTFILE = 'mean'.
MEANS TABLES=condition_score BY condition
/CELLS MEAN SEMEAN.
OMSEND.
dataset activate mean.
compute mean_minus = mean - Std.ErrorofMean.
compute mean_plus = mean + Std.ErrorofMean.
execute.
select if Var1 "Total".
execute.
rename variables (Var1 = condition).
*Example just interval bars.
GGRAPH
/GRAPHDATASET dataset = mean NAME="graphdataset2" VARIABLES=condition mean_plus
mean_minus Mean[LEVEL=SCALE]
MISSING=LISTWISE REPORTMISSING=NO
/GRAPHSPEC SOURCE=INLINE.
BEGIN GPL
SOURCE: s2=userSource(id("graphdataset2"))
DATA: condition=col(source(s2), name("condition"), unit.category())
DATA: mean_plus=col(source(s2), name("mean_plus"))
DATA: mean_minus=col(source(s2), name("mean_minus"))
DATA: Mean=col(source(s2), name("Mean"))
GUIDE: axis(dim(1), label("Var1"))
GUIDE: axis(dim(2), label("Mean Estimate and Std. Error of Mean"))
SCALE: linear(dim(2), include(0))
ELEMENT: interval(position(region.spread.range(condition*(mean_minus+mean_plus))),
shape(shape.ibeam))
ELEMENT: point(position(condition*Mean), shape(shape.square))
END GPL.
*now to put the two datasets together in one chart.
*note you need to put the dynamite source first, otherwise it treats it as a dataset with one observation!
*also needed to do some post-hoc editing to get the legend to look correct, what I did was put an empty text box over top of
*the legend items I did not need.
GGRAPH
/GRAPHDATASET dataset = mean NAME="graphdataset2" VARIABLES=condition mean_plus
mean_minus Mean[LEVEL=SCALE]
MISSING=LISTWISE REPORTMISSING=NO
/GRAPHDATASET dataset = dynamite NAME="graphdataset" VARIABLES=condition condition_score MISSING=LISTWISE
REPORTMISSING=NO
/GRAPHSPEC SOURCE=INLINE.
BEGIN GPL
SOURCE: s=userSource(id("graphdataset"))
DATA: condition2=col(source(s), name("condition"), unit.category())
DATA: condition_score=col(source(s), name("condition_score"))
SOURCE: s2=userSource(id("graphdataset2"))
DATA: condition=col(source(s2), name("condition"), unit.category())
DATA: mean_plus=col(source(s2), name("mean_plus"))
DATA: mean_minus=col(source(s2), name("mean_minus"))
DATA: Mean=col(source(s2), name("Mean"))
GUIDE: axis(dim(1), label("Condition"))
GUIDE: axis(dim(2), label("Tendency Score"))
SCALE: linear(dim(2), include(0))
SCALE: cat(aesthetic(aesthetic.color.interior), map(("Observation", color.grey), ("Mean", color.black), ("S.E. of Mean", color.black)))
SCALE: cat(aesthetic(aesthetic.color.exterior), map(("Observation", color.grey), ("Mean", color.black), ("S.E. of Mean", color.black)))
SCALE: cat(aesthetic(aesthetic.shape), map(("Observation", shape.circle), ("Mean", shape.square), ("S.E. of Mean", shape.ibeam)))
ELEMENT: point.dodge.symmetric(position(condition2*condition_score), shape("Observation"), color.interior("Observation"), color.exterior("Observation"))
ELEMENT: interval(position(region.spread.range(condition*(mean_minus+mean_plus))),
shape("S.E. of Mean"), color.interior("S.E. of Mean"), color.exterior("S.E. of Mean"))
ELEMENT: point(position(condition*Mean), shape("Mean"), color.interior("Mean"), color.exterior("Mean"))
END GPL.
*******************************************************************************************.R code using ggplot2 to generate dot plot
library(ggplot2)
library(reshape)
#this is where I saved the associated dat file in the post
work <- "F:\\Forum_Post_Stuff\\dynamite_plot"
setwd(work)
#reading the dat file provided in question
score <- read.table(file = "exp2tend.dat",header = TRUE)
#reshaping so different conditions are factors
score_long <- melt(score)
#now making base dot plot
plot <- ggplot(data=score_long)+
layer(geom = 'point', position =position_dodge(width=0.2), mapping = aes(x = variable, y = value)) +
theme_bw()
#now making the error bar plot to superimpose, I'm too lazy to write my own function, stealing from webpage listed below
#very good webpage by the way, helpful tutorials in making ggplot2 graphs
#http://wiki.stdout.org/rcookbook/Graphs/Plotting%20means%20and%20error%20bars%20(ggplot2)/
##################################################################################
## Summarizes data.
## Gives count, mean, standard deviation, standard error of the mean, and confidence interval (default 95%).
## data: a data frame.
## measurevar: the name of a column that contains the variable to be summariezed
## groupvars: a vector containing names of columns that contain grouping variables
## na.rm: a boolean that indicates whether to ignore NA's
## conf.interval: the percent range of the confidence interval (default is 95%)
summarySE <- function(data=NULL, measurevar, groupvars=NULL, na.rm=FALSE, conf.interval=.95, .drop=TRUE) {
require(plyr)
# New version of length which can handle NA's: if na.rm==T, don't count them
length2 <- function (x, na.rm=FALSE) {
if (na.rm) sum(!is.na(x))
else length(x)
}
# This is does the summary; it's not easy to understand...
datac <- ddply(data, groupvars, .drop=.drop,
.fun= function(xx, col, na.rm) {
c( N = length2(xx[,col], na.rm=na.rm),
mean = mean (xx[,col], na.rm=na.rm),
sd = sd (xx[,col], na.rm=na.rm)
)
},
measurevar,
na.rm
)
# Rename the "mean" column
datac <- rename(datac, c("mean"=measurevar))
datac$se <- datac$sd / sqrt(datac$N) # Calculate standard error of the mean
# Confidence interval multiplier for standard error
# Calculate t-statistic for confidence interval:
# e.g., if conf.interval is .95, use .975 (above/below), and use df=N-1
ciMult <- qt(conf.interval/2 + .5, datac$N-1)
datac$ci <- datac$se * ciMult
return(datac)
}
##################################################################################
summary_score <- summarySE(score_long,measurevar="value",groupvars="variable")
ggplot(data = summary_score) +
layer(geom = 'point', mapping = aes(x = variable, y = value)) +
layer(geom = 'errorbar', mapping = aes(x = variable, ymin=value-se,ymax=value+se))
#now I need to merge these two dataframes together and plot them over each other
#merging summary_score to score_long by variable
all <- merge(score_long,summary_score,by="variable")
#adding variables to data frame for mapping aesthetics in legend
all$observation <- "observation"
all$mean <- "mean"
all$se_mean <- "S.E. of mean"
#these define the mapping of categories to aesthetics
cols <- c("S.E. of mean" = "black")
shape <- c("observation" = 1)
plot <- ggplot(data=all) +
layer(geom = 'jitter', position=position_jitter(width=0.2, height = 0), mapping = aes(x = variable, y = value.x, shape = observation)) +
layer(geom = 'point', mapping = aes(x = variable, y = value.y, color = se_mean)) +
layer(geom = 'errorbar', mapping = aes(x = variable, ymin=value.y-se,ymax=value.y+se, color = se_mean)) +
scale_colour_manual(" ",values = cols) +
scale_shape_manual(" ",values = shape) +
ylab("[pVisual - pAuditory]") + xlab("Condition") + theme_bw()
plot
#I just saved this in GUI to png, saving with ggsave wasn't looking as nice
#changing width/height in ggsave seems very strange, maybe has to do with ymax not defined?
#ggsave(file = "Avoid_dynamite.png", width = 3, height = 2.5)
#adjusting size of plot within GUI works just fineFeel free to let me know of any suggested improvements in the code. The reason I did code both in SPSS and R is that I was unable to generate a suitable legend in SPSS originally. I was able to figure out how to generate a legend in SPSS, but it still requires some post-hoc editing to eliminate the extra aesthetic categories. Although the chart is simple enough maybe a legend isn’t needed anyway.