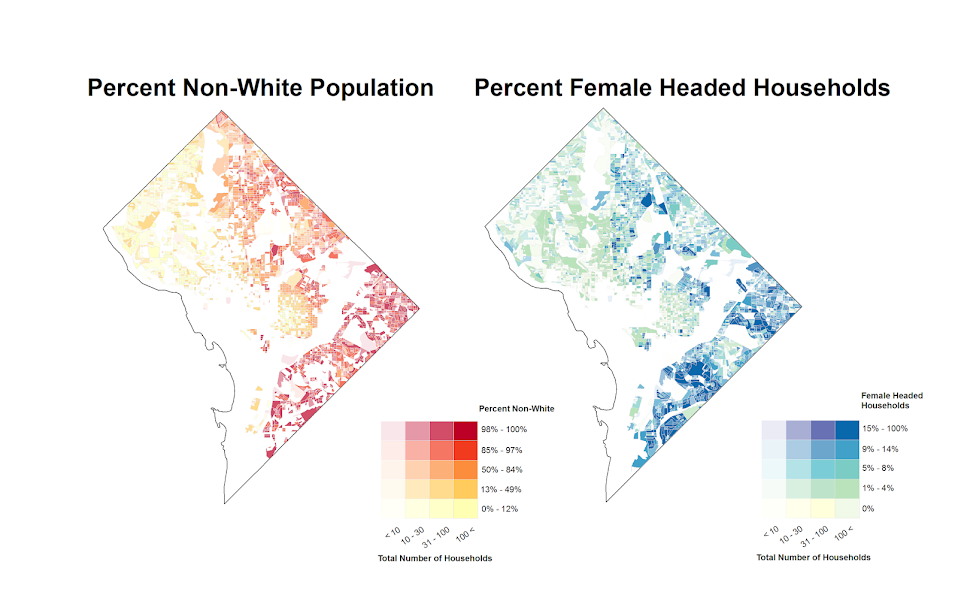
Here is a problem I’ve encountered a few times in my own work (and others) with Poisson regression models and the exponential link function. It came up recently in some discussions on the scatterplot blog by Jeremy Freese (see 1 & 2) critiquing the PNAS paper on the effect of female named hurricanes on death tolls, so I figured I would expand up those thoughts a little here.
So the problem is when you estimate a Poisson regression model is that the exponential link function can become explosive for explanatory variables that have a large range. So to be clear, we have a Poisson regression model of the form (here E[Y] mean the expected value of Y):
log(E[Y]) = B1*(X)
E[Y] = e^(B1*X)If X has a small range this may be fine, but if X has a large range it can become problematic. Consider if Y are hurricane deaths, X is monetary damage of the hurricane, and B1 = 0.01. Lets say the monetary damage ranges from 1 to 1000 (imagine these are in thousands of dollars, so range between $1,000 and $1 million). What happens with the predictions?
E[Deaths] = e^(0.01* 1) = 1.01
E[Deaths] = e^(0.01* 5) = 1.05
E[Deaths] = e^(0.01* 10) = 1.11
E[Deaths] = e^(0.01* 50) = 1.65
E[Deaths] = e^(0.01* 100) = 2.71
E[Deaths] = e^(0.01* 500) = 148
E[Deaths] = e^(0.01*1000) = 22026These predictions are invariant to linear transformations – that is Z-scoring X in the original units doesn’t change the predictions (the same as expressing X in [dollars/1000] doesn’t make any difference than just by including [X] on the right hand side). The linear predictor of B1 will simply be scaled by the appropriate inverse transformation. Also I’d note that expressed in terms of incident rate ratios the effect would be e^0.01=1.01. This appears on its face a totally innocuous effect, and only in consideration of the variation in X does it appear to be absurd.
You can see that if the range of X were smaller, say between 1 and 100, the predictions might be fine. The predictions between 1 and 100 only vary by 1.7 deaths. The problem with these explosive predictions at larger values is that they are nonsense for most of social scientific research. A simple sanity check to see if this is occurring is to check the predicted value from your Poisson regression equation at the low end of X versus the high end (and just pretend all of the other explanatory variables are set to 0) exactly as I have done here. If the high end is crazy, you will need to consider some alternative model specification (or be very clear that the model can not be extrapolated to the larger values of X).
A useful alternate parametrization is simply to log X, and in this case when exponentiating the right hand side, it will make the predictor a power of the original metric.
So imagine we fit the model:
log(E[Y]) = B2*(log(X))
E[Y]) = e^(B2*log(X))
= x^B2Lets say here that B2 = 0.5. What happens to our predictions again?
E[Deaths] = 1^0.5 = 1
E[Deaths] = 5^0.5 = 2.2
E[Deaths] = 10^0.5 = 3.2
E[Deaths] = 50^0.5 = 7.1
E[Deaths] = 100^0.5 = 10
E[Deaths] = 500^0.5 = 22
E[Deaths] = 1000^0.5 = 32Those predictions look a little bit easier to swallow at the larger ranges. Notice also the differences in predictions in the smaller stages? There is more discrimination for the smaller values than on the original scale, but the larger values are suppressed. Lets consider the predictions side by side for easier comparison.
X B1 B2
- -- --
1 1 1
5 1 2
10 1 3
50 2 7
100 3 10
500 148 22
1000 22026 32A frequent problem with logging the explanatory variables is that they contain zeroes. A simple alternative is to treat log(0) as 0 and then have a separate dummy variable equal to 1 when X = 0. This model may not make Occam happy, as it implies a discontinuity at 0, but it is in my opinion a small price to pay. Also if there are a lot of zeroes this doesn’t strike me as totally unrealistic to have a mixture of what happens at 0 and then what happens at the higher values. So the full model written out would be:
log(E[Y]) = B3*D + B4*(log(X))But the model is essentially discontinuous. When X=0, we treat log(X)=0 and D=1, so the model reduces to;
log(E[Y]) = B3*D :When X = 0When X>0, D=0 and the model reduces to:
log(E[Y]) = B4*(log(X)) :When X > 0Now, it certainly would be weird if B3>>0, as this would imply a high spike at 0, and then at the X value of 1 Y goes back down to 1 and then increases with X. If we expect B4 to be positive, then a negative value of B3 (or very close to 0) would make the most sense. It is still a discontinuity in the function, but one that may make theoretical sense. So imagine we fit the equation log(E[Y]) = B3*D + B4*(log(X)), lets say B4 is 0.5 (the same as B2), and that B3 is equal to -0.1. This would then make the set of predictions go:
E[Deaths] = e^-0.01 = 0.9
E[Deaths] = 1^0.5 = 1.0
E[Deaths] = 5^0.5 = 2.2
E[Deaths] = 10^0.5 = 3.2
E[Deaths] = 50^0.5 = 7.1
E[Deaths] = 100^0.5 = 10
E[Deaths] = 500^0.5 = 22
E[Deaths] = 1000^0.5 = 32So in this made up example the discontinuity pretty much fits right in with the rest of the function. We may consider other non-linear transformations of X as well (splines or higher powers) but frequently an additional problem is that the bulk of the data lie in the lower end of the range. So for our dollars if it was highly right skewed, there may only be a few values at 100 or higher. These can be highly influential if you use powers of X (e.g. include X^2, X^3 etc. on the right hand side) – so splines are a better choice – but essentially no matter how you fit the function it will be hard to verify the fit at these values or extrapolate to those tails. So the fit in the original function may be fine – but it just implies unrealistic marginal effects in the tails.
So how do we verify one equation over the other? Visualizing count data in scatterplots tend to be harder than visualizing continuous data, especially if there is a stock pile of data at 0. The problem is simply exacerbated if the explanatory variable has a similar right skew, there will be a large mass near the origin of the plot and very sparse everywhere else.
My simple suggesting is to just bin the data at X values, which is very simple if X is integer valued, and then plot the mean and standard error of the Y value within those bins. As Poisson regression and its variants rely on asymptotic properties, if the error bars are too variable to deduce a pattern you should be concerned your sample size isn’t large enough to begin with.
If the bulk of the data only have a small range over X, then it will be hard in practice to differentiate between the two model parametrizations I suggest here (with the typical noisy data we have in the social sciences). So you may prefer logging the X variable simply to prevent the dramatic explosions in the tails of the data right from the start.
I do feel comfortable saying that if the ratio of the smallest to largest value for the independent variable is over 100, you should check the predictions of the exponential link function very closely (if the smallest value is 0 just estimate the ratio as if the smallest value is 1). If this ratio is 100 or larger, unless the linear predictor in the Poisson regression equation is very small (<<.01) the predictions may explode into very implausible ranges for the larger X values.